Salmonella on xld1/14/2024  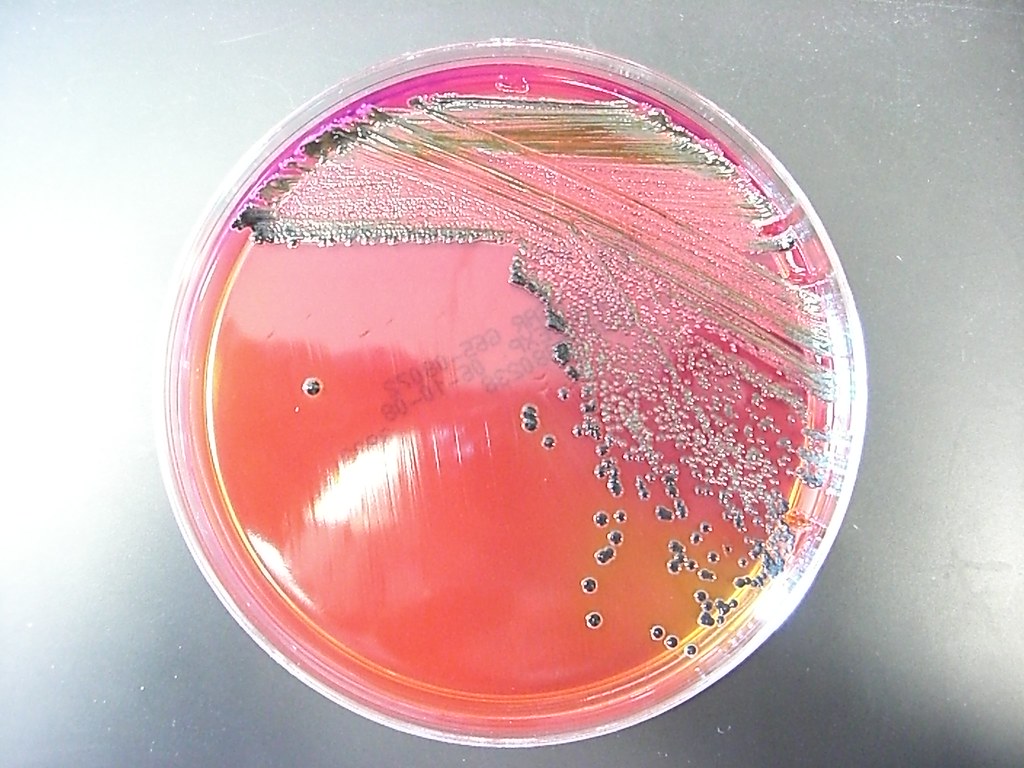
the color disappears after 24 hours, so observations should be made between 18 and 24 hours ⚡ Incubation for more than 48 hours may lead to false positive results. coli Partial inhibition Yellow colonies Enterobacter / Klebsiella Partial inhibition Yellow colonies / Yellow and mucoid colonies Proteus Red to yellow colonies, some strains of Proteus will give colonies with a black center Pseudomonas Partial inhibition Red colonies "Shigella-like" Enterococcus Inhibited / Gram-positive bacteria Inhibited / et Salmonella H2S negative, Providencia Good Red colonies E. ⚡ The inclusion of an H2S indicator system composed of sodium thiosulphate and ferric ammonium citrate improves the differentiation capacity of the preparation: this makes it possible to visualize the production of hydrogen sulphide, which causes the formation of colonies with black centers.īacteria Growth Results Salmonella H2S positive, Edwardsiella Good Red colonies with black center Shigella spp. ⚡ Yellow colonies are also observed for lysine-negative organisms, such as Proteus species. These organisms include Escherichia, Klebsiella, Serratia, Citrobacter koseri, Yersinia enterocolitica. Other uninhibited organisms capable of fermenting lactose or sucrose will produce yellow colonies due to continued acid production, the high level of acid produced prevents the pH from returning to an alkaline value, and d 'a resulting change in pH. However, the presence of Salmonella and Edwardsiella spp is differentiated from that of shigella by an indicator of hydrogen sulfide. Organisms capable of fermenting xylose only, such as Salmonella, will deplete the xylose supply and start using lysine thereby changing the pH to alkaline and the color returning to red. 
Sodium thiosulfate and ferric ammonium serve as indicators of hydrogen sulfide (H2S) production under alkaline conditionsĮ.coli on XLD unable to utilize carbohydrates, such as Shigella and Edwardsiella, will not produce any significant changes and therefore colonies and medium will remain red after incubation.This alkaline pH brings back the red color of the medium. The decarboxylation of lysine, in the absence of lactose and sucrose fermentation, results in a return to an alkaline pH.Fermentation of sugars brings the pH of the medium to an acidic state and the color of the medium turns yellow Xylose, lactose and sucrose, together with phenol red, are fermentable carbohydrates.◈ Differentiation : Is done by three indicator systems ◈ Selectivity : The selective agent is sodium deoxycholate, a bile salt, which inhibits the growth of gram-positive organisms. The three added carbohydrate sources are present in different concentrations, xylose is limited while lactose and sucrose are considered inexhaustible during the prescribed incubation period. The chart above (partial inhibition) needs quantifying.Xylose lysine deoxycholate (XLD) agar is a selective, differential and indicator medium. Sadly, E.coli I have no experience on XLD. More often been hampered by good growth Citrobacter spp.and mimicing Salmonella, even in O-serology IMEX mainly (questionably ) focussed on + ve controls / multi-media/multi-broths for Salmonella. I have never used quartering so cannot comment much in that respect. I can find refs to defined strains of E.coli, eg. Are you using a defined strain, ie one defined to have minimal growth.? or a possibly contaminated self-defined strain. As per my previous post, this genus/species can be problematic. I cannot find any refs/media specs using defined strains (or sometimes self-defined ) of citrobacter spp as a -ve control. Or, as in yr No.2 of post 6, perhaps E.coli - strain XXXX should replace the C.x above.? I guess from title of this thread - probably YES. Partial to complete inhibition yellow to yellow red coloniesĪttached here are some files of XLD MEDIA-Īs i understand, yr query relates to (a) applying species C.x, strain XXXX to brand Y, XLD plates and (b) getting excessive yellow colonies on the plate (ie >G1) when minimal growth was expected. Partial to complete inhibition clear, pinpoint colonies Good growth red colonies with black centre paratyphi A)Įscherichia, Enterobacter, Klebsiella, Citrobacter, Proteus, Serratia Shigella, Providencia, H 2S-negative Salmonella (e.g. coli is gennerally inhibited and Red colonies may occur with some Proteus and Pseudomonas species. Salmonella,shigella ,e coli and citrobacter are a group of organisms belonging to enterobacteriaceae (coliforms) and citrobacter and e coli are different organisms belonging to different genera.Ĭitrobacter on XLD will give yellow colonies (due to fermentation of xylose) and e. 
0 Comments
Leave a Reply.AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
